Segmentation and Measurement of Spots Segment and Threshold the Spots You can also get some more tips from the ImageJ website. In ROI manager, select Show all (bottom right) to see the boundaries of the cells.Ĭongratulations, you segmented the cells in an unbiased way! If you are not satisfied with the results play around with different thresholding and blurring settings. Now we’ll check how good the segmentation was. Repeat this process for each cell-they will show up in the list in the ROI manager. Then press T on your keyboard-a ROI manager window will appear and the first entry in the list will be your first cell. Use the Wand tool and click on the binary mask of the first cell – you should see a yellow border appearing around it. Next, save individual cells as Region of Interests (ROI). Preparation of Segmented Objects for Analysis The white objects represent individual cells. In the end you should get an image similar to Figure 2.įigure 2: A binary image after Gaussian blur, thresholding and watershedding. To separate these cells into individual objects use the Watershed algorithm by selecting Process->Binary->Watershed. 
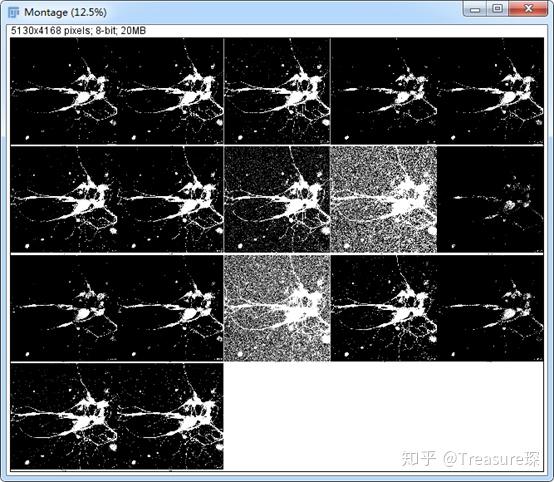
The result is a binary image with one big object representing multiple cells. Don’t forget to tick the Dark background option and then click Auto and Apply. Play with different automated thresholding techniques and see which most closely matches the actual cell area-in our case this is the Mean method. You can then segment by thresholding the image. Apply the Gaussian blur from the menu by selecting Process->Filters. Make a duplicate of the image by right-clicking on the image, select Duplicate and OK. Try the steps below and you’ll quickly see how blurring helps with the segmentation.įirst, open the image in Fiji by dragging it into the main window. The solution is to first blur the image by applying a Gaussian filter with high sigma-values between 10 and 30 pixels usually work fine. Try it yourself by opening the above image and selecting Image->Adjust->Threshold, and then play around with different methods for automatic thresholding. While this works fine if the cell is very bright, it does not work well in our case where we have small bright spots. Because the cells are on a dark background, you could tell Fiji to simply remove everything that is dark. However, a more unbiased approach is to use a method called thresholding. This will be used to tell the computer where the cells are in the image.įor segmentation, you could simply segment the cell out of the background by drawing a polygon or freehand shape selection around it. At the end of this part we’ll have new image with black and white regions-white representing cells and black representing the background. 
We’ll first segment the cells in Figure 1 to demonstrate how it works. But in our case we will stick to the basics. Several image segmentation competitions exist, which demonstrates the complexity and demands of this field. Segmentation includes steps to identify the objects in the image and draw their boundaries. Image segmentation is probably the most important part of the analysis pipeline and the step in which most of the things can go wrong. Fiji can be downloaded from fiji.sc free of charge and is available for Windows, Mac, and Linux. I would recommend that you install Fiji, which is an enhanced distribution of ImageJ and includes multiple image converters and analytical plugins. The above task can be done with the help of ImageJ, which is an open-source image analysis software. For this analysis, we will count how many of these punctates are in a cell, and we will do it in a completely automated/unbiased fashion.įigure 1: An image of several cells with fluorescently stained punctates. They are probably some vesicles that transport material from one compartment to the other. I’ll demonstrate these parts in a few examples.įigure 1 shows an image of a cell, which has fluorescently labeled punctate structures. The analytical pipeline of phenotype quantification is actually quite complex, but can be separated into three main parts: In this article, I’ll give you an introduction to unbiased bio-image analysis and show you some tools that will help you quantify images. Many image analysis approaches exist nowadays, from simple manual measurements to more advanced machine learning models. To remain objective, we try to use analytical approaches to quantify images and turn phenotypes into numbers. The so-called belief in this context is a product of highly experienced imagination, which raises a concern about the objectivity. Microscopists like to say that seeing is believing. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
